Accelerating the Diagnosis and Treatment of Cancer with Mass-Spectrometry Based Techniques
An interview with Livia Eberlin, Assistant Professor at the University of Texas at Austin, discussing the development of mass-spectrometry based techniques for cancer diagnosis and improved clinical outcomes.
Why is there a need for alternatives to cytological analysis in the field of oncology?
Within the context of cancer surgeries, the method that has been in the clinic for analyzing tissues that are being removed from the patient to make sure that all the cancer has been removed is called a histopathology.
That’s done through the analysis of frozen sections. These are pieces of tissue that are sent to a laboratory and then they’re frozen, sectioned, and stained. The pathologist then looks at the stained tissue section under a microscope to determine if all the cancer has been removed or not during that surgical procedure.
The problem is that this procedure can take a long time. Sometimes the surgeries are extended by 35 to 40 minutes because you have to wait for the frozen section analysis to be done. It can also be quite subjective because this process of freezing the tissue quickly during surgery to be able to get a section for the pathologist to look at using a microscope can cause some changes to the tissue histology and the cytology. It can be hard for a pathologist to precisely evaluate if there’s cancer or not in that region of the tissue section.
There’s an incredible need for new technologies that could be applied in the clinic, preferably in the operating room, to help the surgeons by providing them with information on the presence or absence of cancer at certain positions of the body.
This would guide the surgical resection so that they would know, at specific positions when they’re removing the cancer, if all the cancer has been removed or not, and if that’s a region of healthy tissue that can remain or diseased tissue that still needs to be removed.
 Image Credit:Shutterstock/Komsan Loonprom
Image Credit:Shutterstock/Komsan LoonpromHow can mass-spectrometry based techniques be used to improve the diagnosis and monitoring of patients with cancer?
Mass spectrometry is one of the analytical technologies that can provide the highest level of sensitivity and specificity for chemical analysis. With mass spectrometry, we can really dig deep into the molecular composition of complex samples and say, “These are the specific metabolites that are present in this sample. These are the lipids that are present, and the proteins as well”.
With that kind of detailed molecular information, we can then evaluate if a sample, let’s say a clinical sample such as a piece of tissue, is diseased or if it’s healthy tissue in a manner of seconds. That’s because the profiles of these molecules are very characteristic of cells that are healthy or cells that are sick i.e. cells that have cancer.
If we can adapt mass spectrometry technology, which is still mostly a research and development technology that requires relatively complex instrumentation, to translate it to the clinic and put these technologies in the hands of the clinicians, let’s say the surgeons or the pathologists who are the clinical professionals making these decisions, we can empower them with molecular information that can be highly accurate in regards to the diagnosis of a clinical specimen. In that way, we can provide new information that’s currently unavailable to help clinicians make better treatment decisions for their patients.
How do these techniques compare to gene and RNA sequencing?
Mass spectrometry techniques are normally used to analyze molecules such as metabolites, lipids, and proteins. In particular, for the technology that I’ve been developing, we’ve mainly focused on small molecules, which would be metabolites, fatty acids, and lipids. When you compare to DNA and RNA-seq, you’re looking at completely different types of molecules. Also, what I think is appealing about metabolic information is that it’s providing a real-time picture of the processes that are going on in the tissue.
With DNA and RNA, you’re looking at the overall mutations or expression patterns of genes and the transcription processes that are going on in cells. With metabolites, you are looking at the end products of these reactions, things that are happening in real-time in the cell related to metabolism, and so on. Both approaches have incredible value.
It is remarkable how DNA and RNA sequencing technologies are increasingly being incorporated within the clinic. It can provide information on the likelihood that someone will develop, let’s say, a certain type of breast cancer, and that has incredible value in managing patients.
With mass spectrometry, what we’re trying to do is to help improve clinical decisions in nearly real-time. Especially with the type of mass spectrometry that my lab does, the analysis that we can provide is completed in a few seconds. If you compare this with DNA and RNA sequencing, that normally takes a longer time. With mass spec, we could be providing this information with high throughput to hopefully expedite and improve the treatment decision for the patient.
Please can you tell us about your recent research into thyroid neoplasia?
Our research with thyroid cancer is focused on helping patients that come into the clinic with a thyroid nodule to know if the nodule is a cancerous nodule that needs to be removed, or if it’s a benign nodule that does not necessarily require surgery.
Thyroid cancer incidence is rapidly increasing in the USA and around the world, and ~50% of people by the age of 60 will find a nodule in their thyroid. The good news is that the majority of thyroid nodules are benign, so these are not cancerous lesions that need to be removed.
Now the problem is that current cytology methods that are used to evaluate cells from a thyroid biopsy under a microscope are often inconclusive. It can be hard for a clinician to say if a nodule is benign or malignant. So a lot of the time, patients go into surgery without even knowing if they have cancer. You can determine if it is cancer during surgery, but in the majority of the times for the follicular type of neoplasms the patient does not have cancer, so the surgery was likely unnecessary. ,.
What we’re trying to do is use mass spectrometry before surgery to analyze these cells from a minimally invasive biopsy of the nodule to accurately determine if the individual does have cancer, and thus surgery, or if the nodule is benign and thus help prevent unnecessary surgeries, which are terrible for the patient, but are also costly and a burden to the healthcare system.
 Image Credit:Shutterstock/ jovan vitanovski
Image Credit:Shutterstock/ jovan vitanovskiWhat were the conclusions of this study, and were the findings significant?
We started this study in thyroid cancer using banked tissues. These are tissues that have been already collected from patients and they’re available as resources for researchers. We analyzed 178 tissues and acquired over a hundred thousand mass spectra. It was a really large data set. We used that information to build statistical classifiers that could determine if a nodule was cancer or if it was just a benign tumor. We tested that and we did well for a specific type of thyroid cancer, papillary thyroid carcinoma. We had over 90% accuracy. For follicular types, we had 83% accuracy, which is really good considering that this type of thyroid nodule cannot be determined with cytology alone.
Then we started a prospective trial study in the clinic, where patients were coming in for a routine biopsy and we got an additional biopsy for our study. We’ve done that with over a hundred patients so far, and we have kept our accuracy in diagnosis at just about 90%.
These results are really exciting because, in a lot of these cases, we had information that could have prevented patients from going into a potentially unnecessary surgery. But we need to do a much larger validation study which we’re planning to do as a multi-center project to validate these findings and prove its value for patient care.
You also developed a ‘MassSpec Pen’ to improve the accuracy of diagnoses. What is this device, and how does it work?
The vision in developing the mass spec pen was to provide a handheld, easy to use device based on mass spectrometry analysis that could be routinely used by surgeons and pathologists. We wanted it to be something that they could handle, to empower them to do the analysis and take advantage of the high accuracy and sensitivity of mass spectrometry analysis in helping them make clinical decisions.
I look at the MasSpec Pen device, it looks pretty simple, and that was the intention. It’s a handheld tool, and although we called it a pen, of course, it doesn’t function as pen, but it works by providing a single droplet of solvent to extract molecules from a tissue. Most of the time, we use water for the pen tip, and we’ve automated the process so that once you touch the tissue and trigger the device with a foot paddle, everything happens without any more user input.
Water is an incredible solvent. Once that water droplet interacts with the tissue, it extracts metabolites, lipids, and even small proteins from the tissue. Then we have a tubing system that transfers that droplet all the way to the mass spectrometer.
Using this setup, we get molecular analysis of these molecules in nearly real-time Based on the pattern of these molecules, we can then tell the surgeon or the clinician using the device, this place where you analyzed this tissue region is cancer or normal tissue. We do that in a timeframe of about 10-15 seconds. The time can be a little shorter or longer mostly depending on how far your mass spectrometer is from the tissue site and the tubing system that we use. We’ve looked at over a thousand tissues in the lab and our accuracies for cancer diagnosis are pretty exciting, around 96%. We’ve been in the clinic testing this device in the operating room in vivo and on freshly excised tissues with over 100 surgeries now, and the intraoperative results are really promising as well
We are in the process of publishing this pilot study, which really shows that mass spectrometry technology has incredible value in guiding clinical decisions. With the mass spec pen, I think the simplicity and ease of use in the way that we designed the technology is really appealing. It can be well incorporated into a clinical workflow with minimal training requirements.
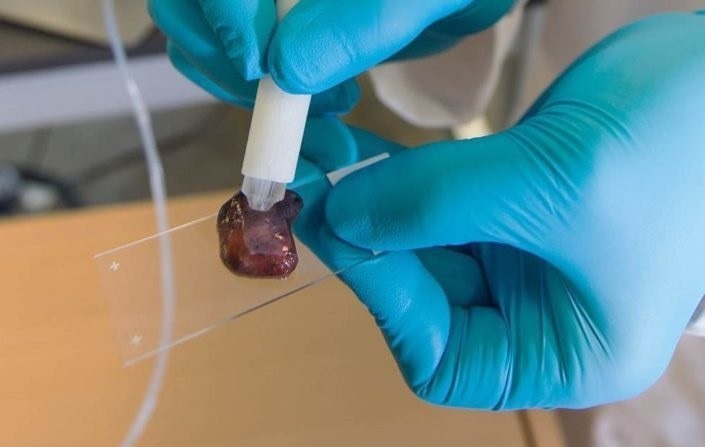 Image Credit: Mass Spec Pen from Livia S. Eberlin Research Group
Image Credit: Mass Spec Pen from Livia S. Eberlin Research GroupWhy did you choose to focus on ovarian cancer?
We have been working on ovarian cancer in my lab for a few years since I started my laboratory, and the focus on ovarian cancer was driven by our desire to help patients and women that are suffering from this disease.
We also had an ongoing project looking at the various aspects of clinical diagnosis of ovarian cancer and treatment outcomes. When we developed the mass spec pen, we knew that in ovarian cancer surgery it can be really hard to identify regions of metastasis, which is where the ovarian cancer is spreading, which is very common for high grade ovarian cancer.
You normally find ovarian cancer throughout the abdominal cavity of a patient. For a surgeon, it can be really important to have a tool that will help them to identify these potential regions of metastasis in order to remove all of the cancer.
We know from clinical data that there have been extensive studies that show that removing all of the cancer from the patient will give them a higher chance of disease-free survival. It was a scenario where we had access to these patients and tissues and there was an important clinical need. We are extremely passionate about helping ovarian cancer patients.
 Image Credit:Shutterstock/David A. Litman
Image Credit:Shutterstock/David A. LitmanThe technique involves the use of machine learning. Do you think AI and machine learning will form a vital part of the future for clinical diagnostics?
I believe that AI and machine learning will be essential parts of the clinical decision, and that will happen sooner than we expect. They are essential technologies, especially as we are constantly moving towards big data and incorporating molecular data with clinical and imaging information.
Once you start to get to these complex data sets and try to make decisions based on this complex information, there’s a limit to what the human brain can do rapidly and automatically. Incorporating AI and machine learning will be critical to achieve that level of throughput and certainty.
But I have to say that there’s a lot of hype in this field as well. It’s really important for researchers like myself, and I’m constantly trying to learn more about AI and machine learning, to select the right tools to make sure that our models are not over-fitting.
Validating your models, testing your models, and the use of independent data sets and clinical samples will be crucial to show the value and the robustness of this type of technology to help clinical care.
Do you think that these techniques will one day overtake histopathology, or work alongside it?
I think that new technologies like mass spectrometry and other modalities will be complementary to what pathologists are already doing. The human aspect of the evaluation of tissue for diagnosis is crucial to patient care.
The thought, or at least my goal, is to empower clinicians with newer technologies that can help them to make decisions. Not to replace them, but to help them to make more informed and better decisions, especially based on the molecular information that is reliable and can be highly predictive of the disease state.
Why did you feel it was important to share your work at Pittcon 2020?
I’ve been to Pittcon several times. It’s a conference that I enjoy very much because it combines several areas of chemistry and analytical chemistry. There is this cool edge and aspect to it, which is all the new technologies that is launched during the conference every year.
I think that seeing the scientific talks and getting this new information about what people are working on and developing, is really exciting and motivates me to also pursue new areas of research. The exposition is also really cool.
Being able to see the vendors and the new products that are being launched to help researchers in academia and industry is something that is special and unique to Pittcon. It is great avenue for networking as well.
Events like Pittcon are really important for me to talk to other scientists and researchers, and to share ideas and see if they have any input. How can we better develop what we’re doing? How can we think of new ways to reach our goals and help patients?
That interaction with other people who are leading in the field is very important for researchers and for students as well. Again, seeing the technology and the products that are being launched by the industry in the companies is also something that is appealing and can help our research and instrumentation.
Events like Pittcon are really important for me to network, to talk to other scientists and other researchers, and to share ideas and see if they have any input. How can we better develop what we’re doing? How can we think of new ways to reach our goals and help patients?
That interaction with people who are leading in the field is very important for researchers and for grad students as well. Again, seeing the technology and the products that are being launched by the industry in the companies is also something that is appealing and can help our research and instrumentation.
About Livia Schiavinato Eberlin – Assistant Professor at the University of Texas at Austin
Livia Schiavinato Eberlin was born and raised in Campinas, São Paulo, Brazil. Her passion for mass spectrometry (MS) started as an undergraduate research assistant at the Thomson Laboratory of the State University of Campinas (UNICAMP).
She received her B.S. in Chemistry from UNICAMP in 2007 and then moved to the USA in 2008 to start a PhD program in Analytical Chemistry at Purdue University under the mentorship of Prof. R. Graham Cooks.
During her PhD, Livia developed and applied ambient ionization MS imaging to human cancer diagnosis. In recognition of her innovative PhD work, Livia received many awards including the Nobel Laureate Signature Award from the American Chemical Society.
In 2012, she started her postdoctoral work at Stanford University under the guidance of Prof. Richard N. Zare, where she continued to develop MS technology for biomedical research. During that time, she received the L’Oréal for Women in Science Fellowship, a K99 pathway to independence award from the NIH/NCI and was listed in the Forbes 30 under 30 list in Science and Healthcare.
In 2016, Livia started her independent career as an Assistant Professor in the Chemistry Department at the University of Texas at Austin. Since then, Livia and her group have received several recognitions for their research.


Comments are closed.